2.5.3. Linear System Solvers¶
sparse matrix/eigenvalue problem solvers live in
scipy.sparse.linalg- the submodules:
dsolve: direct factorization methods for solving linear systemsisolve: iterative methods for solving linear systemseigen: sparse eigenvalue problem solvers
all solvers are accessible from:
>>> import scipy.sparse.linalg as spla >>> spla.__all__ ['LinearOperator', 'Tester', 'arpack', 'aslinearoperator', 'bicg', 'bicgstab', 'cg', 'cgs', 'csc_matrix', 'csr_matrix', 'dsolve', 'eigen', 'eigen_symmetric', 'factorized', 'gmres', 'interface', 'isolve', 'iterative', 'lgmres', 'linsolve', 'lobpcg', 'lsqr', 'minres', 'np', 'qmr', 'speigs', 'spilu', 'splu', 'spsolve', 'svd', 'test', 'umfpack', 'use_solver', 'utils', 'warnings']
2.5.3.1. Sparse Direct Solvers¶
- default solver: SuperLU 4.0
- included in SciPy
- real and complex systems
- both single and double precision
- optional: umfpack
- real and complex systems
- double precision only
- recommended for performance
- wrappers now live in
scikits.umfpack - check-out the new
scikits.suitesparseby Nathaniel Smith
Examples¶
import the whole module, and see its docstring:
>>> from scipy.sparse.linalg import dsolve >>> help(dsolve)
both superlu and umfpack can be used (if the latter is installed) as follows:
prepare a linear system:
>>> import numpy as np >>> from scipy import sparse >>> mtx = sparse.spdiags([[1, 2, 3, 4, 5], [6, 5, 8, 9, 10]], [0, 1], 5, 5) >>> mtx.todense() matrix([[ 1, 5, 0, 0, 0], [ 0, 2, 8, 0, 0], [ 0, 0, 3, 9, 0], [ 0, 0, 0, 4, 10], [ 0, 0, 0, 0, 5]]) >>> rhs = np.array([1, 2, 3, 4, 5], dtype=np.float32)
solve as single precision real:
>>> mtx1 = mtx.astype(np.float32) >>> x = dsolve.spsolve(mtx1, rhs, use_umfpack=False) >>> print(x) [106. -21. 5.5 -1.5 1. ] >>> print("Error: %s" % (mtx1 * x - rhs)) Error: [0. 0. 0. 0. 0.]
solve as double precision real:
>>> mtx2 = mtx.astype(np.float64) >>> x = dsolve.spsolve(mtx2, rhs, use_umfpack=True) >>> print(x) [106. -21. 5.5 -1.5 1. ] >>> print("Error: %s" % (mtx2 * x - rhs)) Error: [0. 0. 0. 0. 0.]
solve as single precision complex:
>>> mtx1 = mtx.astype(np.complex64) >>> x = dsolve.spsolve(mtx1, rhs, use_umfpack=False) >>> print(x) [106. +0.j -21. +0.j 5.5+0.j -1.5+0.j 1. +0.j] >>> print("Error: %s" % (mtx1 * x - rhs)) Error: [0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
solve as double precision complex:
>>> mtx2 = mtx.astype(np.complex128) >>> x = dsolve.spsolve(mtx2, rhs, use_umfpack=True) >>> print(x) [106. +0.j -21. +0.j 5.5+0.j -1.5+0.j 1. +0.j] >>> print("Error: %s" % (mtx2 * x - rhs)) Error: [0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
"""
Solve a linear system
=======================
Construct a 1000x1000 lil_matrix and add some values to it, convert it
to CSR format and solve A x = b for x:and solve a linear system with a
direct solver.
"""
import numpy as np
import scipy.sparse as sps
from matplotlib import pyplot as plt
from scipy.sparse.linalg.dsolve import linsolve
rand = np.random.rand
mtx = sps.lil_matrix((1000, 1000), dtype=np.float64)
mtx[0, :100] = rand(100)
mtx[1, 100:200] = mtx[0, :100]
mtx.setdiag(rand(1000))
plt.clf()
plt.spy(mtx, marker='.', markersize=2)
plt.show()
mtx = mtx.tocsr()
rhs = rand(1000)
x = linsolve.spsolve(mtx, rhs)
print('rezidual: %r' % np.linalg.norm(mtx * x - rhs))
2.5.3.2. Iterative Solvers¶
- the
isolvemodule contains the following solvers: bicg(BIConjugate Gradient)bicgstab(BIConjugate Gradient STABilized)cg(Conjugate Gradient) - symmetric positive definite matrices onlycgs(Conjugate Gradient Squared)gmres(Generalized Minimal RESidual)minres(MINimum RESidual)qmr(Quasi-Minimal Residual)
- the
Common Parameters¶
mandatory:
- A : {sparse matrix, dense matrix, LinearOperator}
The N-by-N matrix of the linear system.
- b : {array, matrix}
Right hand side of the linear system. Has shape (N,) or (N,1).
optional:
- x0 : {array, matrix}
Starting guess for the solution.
- tol : float
Relative tolerance to achieve before terminating.
- maxiter : integer
Maximum number of iterations. Iteration will stop after maxiter steps even if the specified tolerance has not been achieved.
- M : {sparse matrix, dense matrix, LinearOperator}
Preconditioner for A. The preconditioner should approximate the inverse of A. Effective preconditioning dramatically improves the rate of convergence, which implies that fewer iterations are needed to reach a given error tolerance.
- callback : function
User-supplied function to call after each iteration. It is called as callback(xk), where xk is the current solution vector.
LinearOperator Class¶
from scipy.sparse.linalg.interface import LinearOperator
- common interface for performing matrix vector products
- useful abstraction that enables using dense and sparse matrices within the solvers, as well as matrix-free solutions
- has shape and matvec() (+ some optional parameters)
- example:
>>> import numpy as np
>>> from scipy.sparse.linalg import LinearOperator
>>> def mv(v):
... return np.array([2*v[0], 3*v[1]])
...
>>> A = LinearOperator((2, 2), matvec=mv)
>>> A
<2x2 _CustomLinearOperator with dtype=float64>
>>> A.matvec(np.ones(2))
array([2., 3.])
>>> A * np.ones(2)
array([2., 3.])
A Few Notes on Preconditioning¶
- problem specific
- often hard to develop
- if not sure, try ILU
- available in
dsolveasspilu()
- available in
2.5.3.3. Eigenvalue Problem Solvers¶
The eigen module¶
arpack* a collection of Fortran77 subroutines designed to solve large scale eigenvalue problemslobpcg(Locally Optimal Block Preconditioned Conjugate Gradient Method) * works very well in combination with PyAMG * example by Nathan Bell:""" Compute eigenvectors and eigenvalues using a preconditioned eigensolver ======================================================================== In this example Smoothed Aggregation (SA) is used to precondition the LOBPCG eigensolver on a two-dimensional Poisson problem with Dirichlet boundary conditions. """ import scipy from scipy.sparse.linalg import lobpcg import matplotlib.pyplot as plt from pyamg import smoothed_aggregation_solver from pyamg.gallery import poisson N = 100 K = 9 A = poisson((N,N), format='csr') # create the AMG hierarchy ml = smoothed_aggregation_solver(A) # initial approximation to the K eigenvectors X = scipy.rand(A.shape[0], K) # preconditioner based on ml M = ml.aspreconditioner() # compute eigenvalues and eigenvectors with LOBPCG W,V = lobpcg(A, X, M=M, tol=1e-8, largest=False) #plot the eigenvectors plt.figure(figsize=(9,9)) for i in range(K): plt.subplot(3, 3, i+1) plt.title('Eigenvector %d' % i) plt.pcolor(V[:,i].reshape(N,N)) plt.axis('equal') plt.axis('off') plt.show()
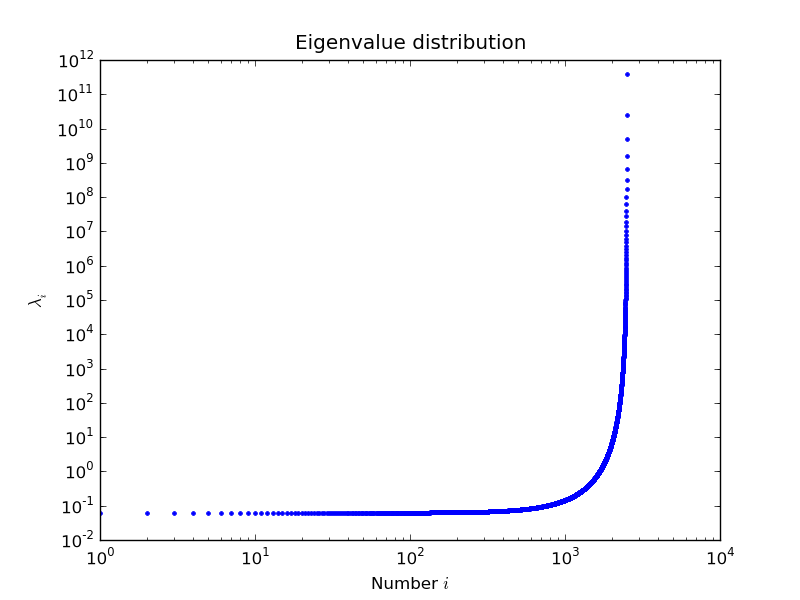
example by Nils Wagner:
output:
$ python examples/lobpcg_sakurai.py Results by LOBPCG for n=2500 [ 0.06250083 0.06250028 0.06250007] Exact eigenvalues [ 0.06250005 0.0625002 0.06250044] Elapsed time 7.01